
Gene editing technologies like transcription activator-like effector nucleases (TALEN), zinc finger nucleases (ZFN), CRISPR-Cas systems, base editors, and prime editors, among others, have become essential tools in a multitude of modern biotechnology applications due to their precise and efficient DNA modification capabilities. These technologies have vast applications in genetic disease treatments, crop improvement, pest control, and bioproduction. They have accelerated research in genetics, molecular biology, and biomedicine. However, a challenge that impedes the speed of scientific advancement using these tools is differences in how the outcome of research efforts is shared and the resulting difficulty researchers then have in ascertaining what work has already been done by other scientists. Science builds upon its own past accomplishments, but only if the knowledge gained can be readily shared and accessed. Towards this end, a specialized digital knowledge base called the Genome Editing Meta-database (GEM) has been created, providing quick access and searchability of past gene editing research efforts.
Though other such databases exist, focusing on published gene editing research, the amount of data they contain is limited. Sometimes being entered into the database manually, seriously hampering the total amount of knowledge that might potentially be gathered. The approach taken to build GEM was different from the outset, relying on automation. To make the approach work, a data source with information categories that could be easily parsed was needed. PubMed/PubMed Central (PMC) served as that bank from which data input could be systematically drawn. Metadata including species, gene of interest and gene editing method used was used to organize the published knowledge, resulting in a database that would be easy to use and sort.
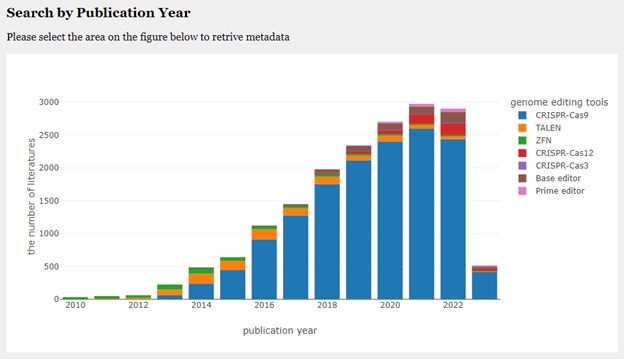
https://bonohu.hiroshima-u.ac.jp/gem/
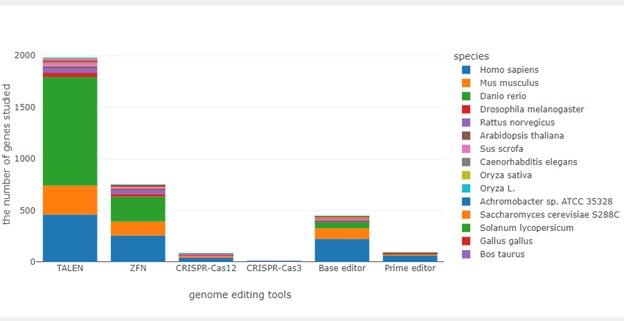
https://bonohu.hiroshima-u.ac.jp/gem/
There are some advantages and disadvantages to the selected data-gathering approach. Being automated makes it possible to repeat the PMC article search on a regular schedule, which is currently set to be monthly. Users of GEM can thus rest assured that information they glean from their GEM searches will never be anymore than a few weeks out-of-date. However, PMC being the data source that the automated system is built around, any scientific articles published elsewhere will be absent from the results.
Nevertheless, the GEM database now enables scientists to efficiently find published results related to a gene or gene pathway their own research may be focused on. They can have greater confidence that their methods reflect the latest scientific best practices, reducing wasted time and resources, as well as helping to prevent inaccurate data that they may have otherwise gone on to publish without realizing it. In addition, the resources they find within GEM might suggest research targets they might not otherwise have been considering, enriching the ultimate outcome of their efforts.
References
Takayuki Suzuki, Hidemasa Bono. GEM: Genome Editing Meta-database, a dataset of genome editing related metadata systematically extracted from PubMed literatures. Gene and Genome Editing, Vol 5, 2023, 100024, ISSN 2666-3880, https://doi.org/10.1016/j.ggedit.2022.100024
