dGH SCREEN™ Assay Services
High-resolution karyotyping assay to monitor genomic stability and structural variants as small as 5kb.
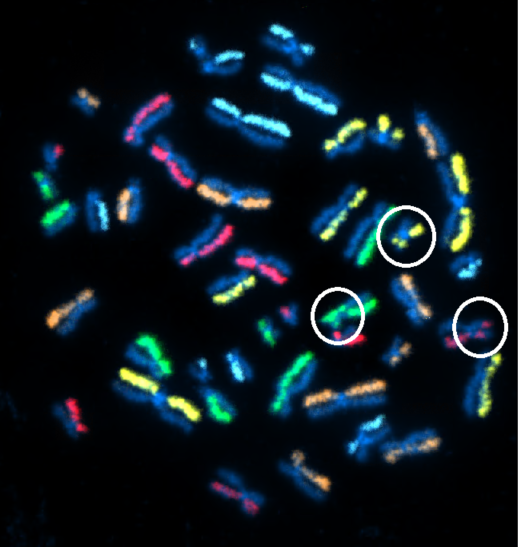
Above Fig.: dGH SCREEN detecting multiple structural rearrangements in metaphase cell from human control cell line.
Map Genomic Structural Variants with dGH SCREEN.
The dGH SCREEN assay provides per-chromosome attribution of inter- and intra-chromosomal structural events including inversions, translocations, aneuploidy (gain and loss), insertions, centromere abnormalities and complex events across a sample. It is a dGH paint-combination assay for all 24 human chromosomes. The assay is composed of unique sequence, high-density (HD) dGH chromosome paints in five color panels such that chromosomes painted in the same color can be differentiated by size, shape, and centromere position.
Use dGH SCREEN to Assess:
- Cellular engineering outcomes: Genome-wide, cell-by-cell and chromosome-by-chromosome structure assessment, pre- and post-modification.
- Structural integrity: Total genomic structural variation metrics; measure the stability of cell lines; screen and compare candidate cell lines.
- Genomic stability: Track persistence of variants over time, passages and process variable changes.
- Hidden variants: Discover unknown mutations by de novo assessment of single cells from patient sub-types.
- Confirm G-banding results: Confirm and characterize large structural rearrangements found by G-banding.
- Sequencing data outcomes: Confirmatory data of rearrangements predicted with long read and NGS analyses.
Request a quote today to discover the power of the dGH SCREEN assay!
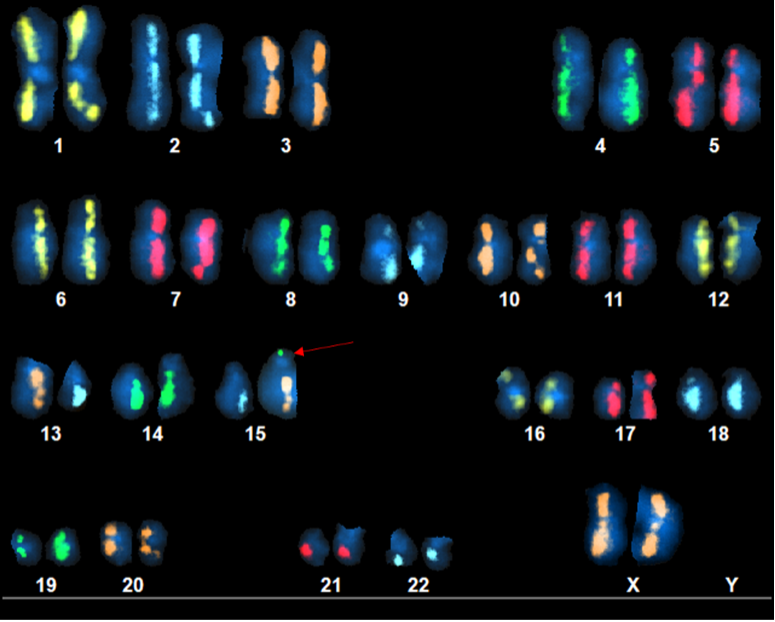
Above Fig.: dGH SCREEN karyogram of a cancer cell line depicting a neoplastic cell revealing a large number of complex rearrangements.
Gene-Edited T cell Evaluated using dGH SCREEN.
CAR-T cell engineering produces outcomes which are difficult to predict. To measure those outcomes you need tools which generate both unbiased and high-resolution data. Aside from a pericentric translocation involving the q-arms of chromosomes 13 and 15 in this karyogram, dGH SCREEN’s fluorescence-based approach also reveals another tiny rearrangement (arrow) which would be very difficult for other assays to detect. G-banding would likely fail to uncover it due to both the small size and location. In addition, the highly repetitive nature of much of the DNA of the acrocentric p-arms would represent a difficult challenge for sequencing platforms. Without any intermediate alignment step, dGH SCREEN easily catches this kind of abnormality.
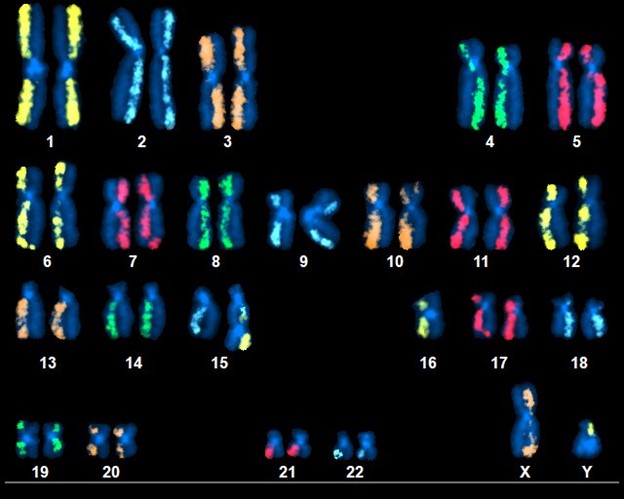
Above Fig.: dGH SCREEN karyogram of a triple-edited T cell displaying multiple inversions and a dicentric derivative composed of material from chromosomes 15 and 16.
dGH SCREEN karyogram of a triple-edited T cell.
Thanks to the unbiased, genome-wide characteristics of dGH SCREEN, detection of chromosomal structural rearrangements, as well as numerical abnormalities is possible without any foreknowledge of which chromosomes or breakpoints may be affected. In this cell, multiple chromosomes display relocation of fluorescence from one chromatid to another, indicating inversions. Also, a dicentric derivative chromosome has resulted from an unbalanced translocation between chromosomes 15 and 16.
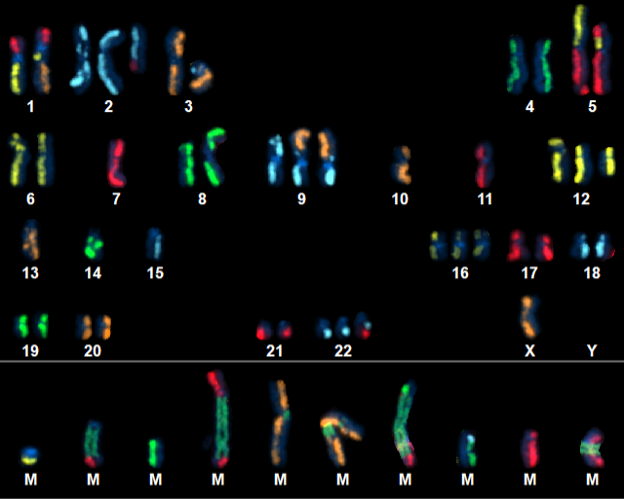
Above Fig.: dGH SCREEN karyogram of a cancer cell line depicting a neoplastic cell revealing a large number of complex rearrangements.
Complex Rearrangements and Inversion Detection.
Generate the highest-resolution, genomic structural variant data for your samples. Comprehensively profile stability and structural variance in your cells – even if they are a complex, heterogeneous population.
dGH SCREEN detects inversions missed by traditional metaphase painting techniques.
Ordering Below
|
Catalog Number |
Services |
Cost |
|---|---|---|
| SCR-003 | 20 spread/sample Assay Execution and Analysis | $3,638 |
| SCR-002 | 50 spreads/sample Assay Execution and Analysis | $6,064 |
| SCR-004 | 100 spreads/sample Assay Execution and Analysis | $10,915 |
| SCR-005 | T-Cell Culture Development: Thaw, Recovery, and Harvest Optimization | $1,788 |
| SCR-009 | T-Cell Metaphase Prep and Harvest | $1,788 |
| SCR-006 | IPSC Cell Culture Development: Thaw, Recovery, and Harvest Optimization | $1,925 |
| SCR-010 | IPSC Metaphase Prep and Harvest | $1,925 |
| SCR-007 | Whole Blood Culture Development: Thaw, Recovery, and Harvest Optimization | $1,375 |
| SCR-011 | Whole Blood Metaphase Prep and Harvest | $1,375 |
| SCR-014 | NK Cell Culture Development: Thaw, Recovery, and Harvest Optimization | $1,788 |
| SCR-013 | NK Cell Metaphase Prep and Harvest | $1,788 |
